CLONE 082514 humKappaCasein(Kex+)
Jump to navigation
Jump to search
Cloning human kappa casein into pD1214 vector, transforming to E. coli, plasmid preps and sequencing
Experiment 082514_humKappaCasein(Kex+) Cloning
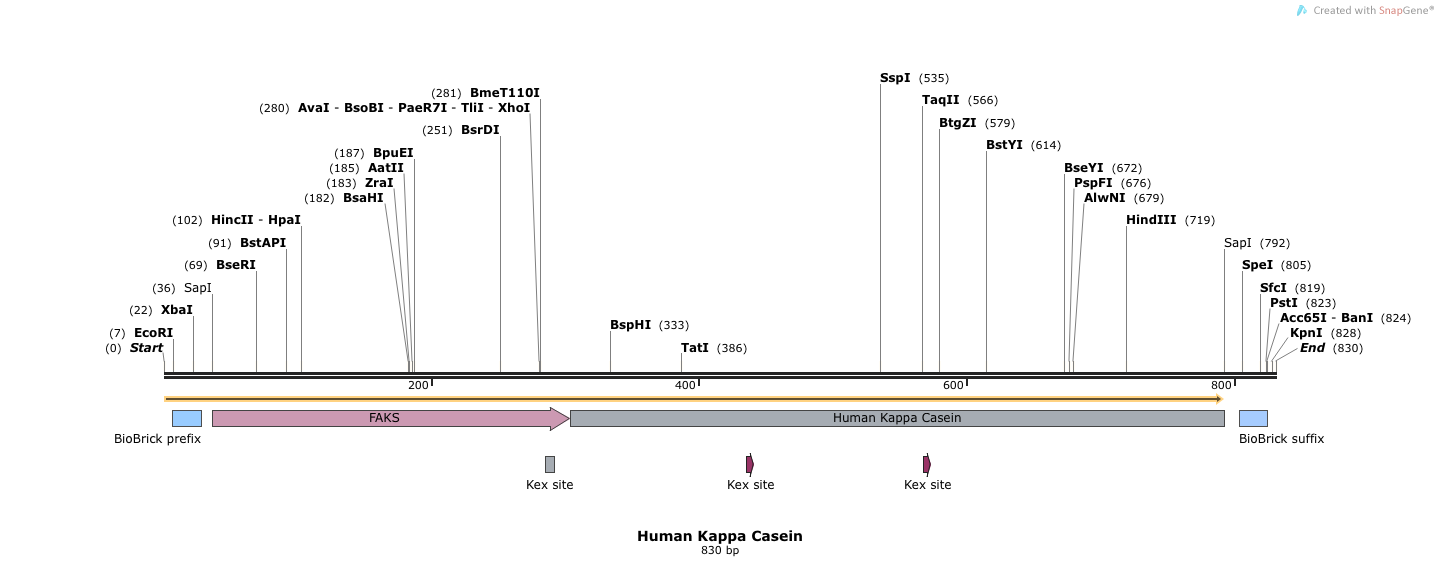
- Human Kappa Casein (BB.SapI.humKappa(Kex+).SapI.BB)
5'- TACACGGAATTCGCGGCCGCTTCTAGAGGCTCTTCTATGAGATTCCCATCTATTTTCACCGCTGTCTTGTTCGCTGCCTCCTCTGCATTGGCTGCCCCTGTTAAC ACTACCACTGAAGACGAGACTGCTCAAATTCCAGCTGAAGCAGTTATCGGTTACTCTGACCTTGAGGGTGATTTCGACGTCGCTGTTTTGCCTTTCTCTAACTCC ACTAACAACGGTTTGTTGTTCATTAACACCACTATCGCTTCCATTGCTGCTAAGGAAGAGGGTGTCTCTCTCGAGAAAAGAGAGGCCGAAGCTGAAGTCCAAAAC CAAAAGCAACCAGCTTGTCATGAAAACGACGAAAGACCATTCTACCAAAAGACTGCCCCATACGTTCCAATGTACTACGTTCCAAACTCTTACCCATACTACGGT ACTAACTTGTACCAAAGAAGACCTGCTATCGCCATTAACAACCCATACGTCCCAAGAACTTACTACGCTAATCCAGCTGTTGTTCGTCCACACGCTCAAATTCCA CAAAGACAATATTTGCCTAACTCTCACCCACCAACCGTTGTCAGAAGACCAAACTTGCATCCTTCTTTCATCGCTATCCCACCAAAAAAGATCCAAGACAAGATT ATCATCCCAACTATCAACACTATCGCCACCGTTGAACCAACCCCAGCTCCTGCCACCGAACCAACTGTCGATTCTGTTGTTACTCCAGAAGCTTTCTCCGAATCT ATCATTACTTCTACTCCAGAAACTACCACTGTCGCCGTCACCCCACCAACTGCTTAGGGTAGAAGAGCTACTAGTAGCGGCCGCTGCAGGTACCA - 3'
- This sequence has 5' and 3' SapI sequences, followed by full length FAKS, followed by human Kappa Casein (Kex+). Electra cloning cleaves at SapI sites and ligates simulatenously.
- We cloned into pD1214. This vector was ordered from DNA2.0 and is linearized upon arrival. Our insert is treated with Electra enzyme mix which cuts and ligates our "SapI.gene.SapI" and inserts it into the pD1214 vector.
- SapI treated DNA has an ATG overhang at the 5' end and a GGT overhang at the 3' end. Our complete vector will have the alphafactorfull signal sequence, which leads into another Methionine (M), followed by the rest of the human kappa casein protein.
All you need to know about the Electra system.
August 25, 2014
- Successfully cloned SapI.hKcasein.SapI into the electra daughter vector (pD1214).
Reaction: 1uL pD1214:FAKS (20ng) 1uL (20ng) of BB.SapI.humKappa(Kex+).SapI.BB 2uL Electra Buffer 1uL of Electra enzyme mix 15uL of Water ---- 20uL, Room Temperature, 20min.
- We stored the cloning reaction at -20 degrees C and waited to transform E. coli cells until August 26, 2014 (Nikola and Johan performed this) since we did not have competent cells available.
Transformation of humKappaCasein(Kex+) in pD1214 to dH5alpha E. coli
August 26, 2014
- Participants: Nikola, Johan
- Location: BioCurious
- Aims: Transform the cloning reaction from August 25, humKappaCasein(Kex+) in pD1214, to chemically competent dH5alpha E. coli
- Materials and Methods: Transforming Mix & Go Competent dH5alpha E. coli with gBlocks in pD1214, above:
- We used the Zymo Research instruction manual for "Premade Mix & Go Competent E. coli Cells." Media:Zymo_E._coli_competent_cells_transformation_protocol.pdf
- We completed 2 reactions: humKappaCasein(Kex+) in pD1214, one pGLO positive control.
- Thawed individual tubes of Mix & Go chemically competent, dH5alpha E. coli cells slowly on ice.
- Prewarmed LB agar plates with Carb-50, 100mm (Teknova), at 37°C.
- Added 5uL of humKappaCasein(Kex+) in pD1214 or pGLO control to individual thawed tube of 100uL competent cells. Mixed gently.
- Immediately incubated on ice for 10 min.
- Plated 100uL of each reaction to LB carb plates, using one individually wrapped spreader/reaction.
- Inducible promoter for pGLO positive control requires arabinose, so 200uL of arabinose spread onto that LB carb plate prior to transformation mix being spread.
- Incubated plates overnight at 37°C.
Midiprep of humKappaCasein(Kex+) in pD1214
August 27, 2014
- Participants: Nikola, Johan, Meenakshi, Lafia, Aaron, Matt
- Location: BioCurious
- Aims: Extract the plasmid DNA from O/N culture of E. coli for further characterization of and experimentation with humKappaCasein(Kex+) in pD1214.
- Materials and Methods: Followed protocol for Zyppy Plasmid Midiprep Kit, Zymo Research: Media:D4025i.pdf.
- Used "Centrifugation Protocol" stream, with the following specific changes to step 1): grew in 12mL of LB Amp and added 6mL of water to the bacterial cell pellet.
DNA quantification of humKappaCasein(Kex+) in pD1214
September 1, 2014
- Participants: Advait, Patrik, Johan, Minakishi, Rachel, Lafia, Mac
- Location: BioCurious
- Aims: Quantify and check purity ratio of humKappaCasein(Kex+) in pD1214 (midiprepped as above, August 28, 2014) in preparation for submitting to Sequetech for sequencing.
- Materials & Methods: All measurements taken on BioCurious Beckman Coulter DU 640 spectrophotometer, DNA/Oligo setting, using 100ul cuvette.
Measuring plasmid DNA diluted 1:100: Blank = 1ul Zyppy kit elution buffer + 99ul diH2O Sample = 1ul plasmid DNA + 99ul diH2O
- Results: Spectrophotometer gave DNA concentration readings of 0. Repeating with a lower dilution factor.
Measuring plasmid DNA diluted 1:10: Blank = 10ul Zyppy kit elution buffer + 90ul diH2O Sample = 10ul plasmid DNA + 90ul diH2O
- Results:
- Read 1: 42.158 ug/mL; 260/280 ratio = 1.9291
- Read 2: 42.433 ug/mL; 260/280 ratio = 1.9382
- Conclusions:
- DNA concentration looks good; 260/280 ratio a bit high ("pure" DNA = 1.8, "pure" RNA = 2.0).
- Construct ready to send to Sequetech for sequencing.
DNA pick-up by Sequetech to sequence humKappaCasein(Kex+) in pD1214 & sequencing results
September 4, 2014
- Location: BioCurious
- Aims: Confirm correct DNA sequence of our alpha factor - human kappa casein insert into pD1214 vector (as mapped, above.)
- Sequencing Strategy & Methods:
- Submitting sample hkckplus, a 10uL aliquot of August 28, 2014 midiprep, to be sequenced in two reactions: the forward (with primer TEF-264-FW) and reverse (with primer PMYR3) directions. Sequetech collecting DNA September 5, 2014 from BioCurious. DNA transport requires a secondary container, so sealed sample microfuge tube inside a 50mL screw-top conical tube.
- TEF-264-FW is a custom primer synthesized through Sequetech: primer sequence recommended by DNA 2.0, suppliers of pD1214. This primer will be maintained in-house for future sequencing reactions. (Note change from sequencing described August 6, 2014, as previous forward primer, Alpha-factor 146-forward, anneals to the FAKS sequence, now inside our redesigned DNA inserts.)
- PMYR3 is an in-house Sequetech primer.
- Submitting sample hkckplus, a 10uL aliquot of August 28, 2014 midiprep, to be sequenced in two reactions: the forward (with primer TEF-264-FW) and reverse (with primer PMYR3) directions. Sequetech collecting DNA September 5, 2014 from BioCurious. DNA transport requires a secondary container, so sealed sample microfuge tube inside a 50mL screw-top conical tube.
Primer Sequences TEF-264-FW: 5’ TCGATGACCTCCCATTGA 3’ Binds to TEF promoter and reads through 5'-3' (N-terminus of signal peptide). PMYR3: 5' CTTCCTTTTCGGTTAGAG 3' Binds to the terminator (CYC1) and reads through in the 3'-5' direction (C-terminus of casein).
Location of sequencing primers indicated by small purple arrows on map of pD1214:
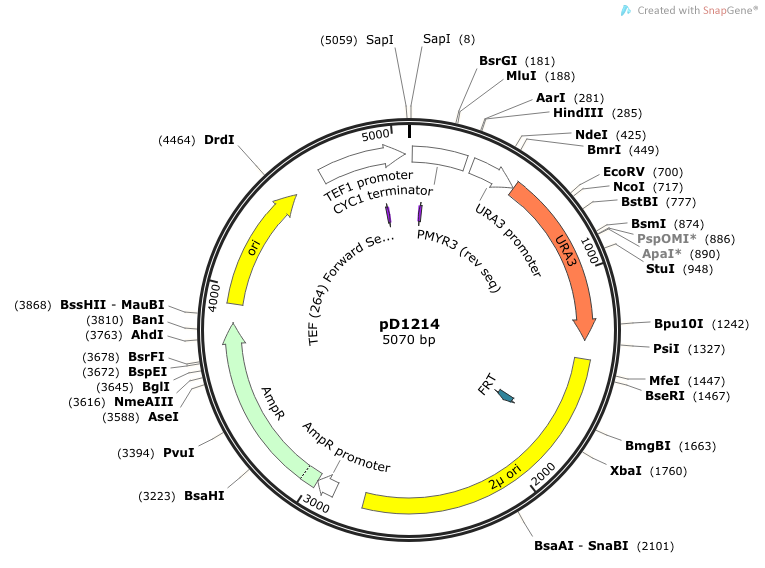
September 5, 2014
- Sequetech collected the humKappaCasein(Kex+) in pD1214 DNA from BioCurious at ~2:00pm today.
September 8, 2014
- Analyzed data and created alignments in SerialCloner Freeware.

- File:HumanKap(Kex+).png