Difference between revisions of "Submitting 10 casein and FAM20C in pD1214 plasmids for sequencing 30Sep2014"
Jump to navigation
Jump to search
| Line 32: | Line 32: | ||
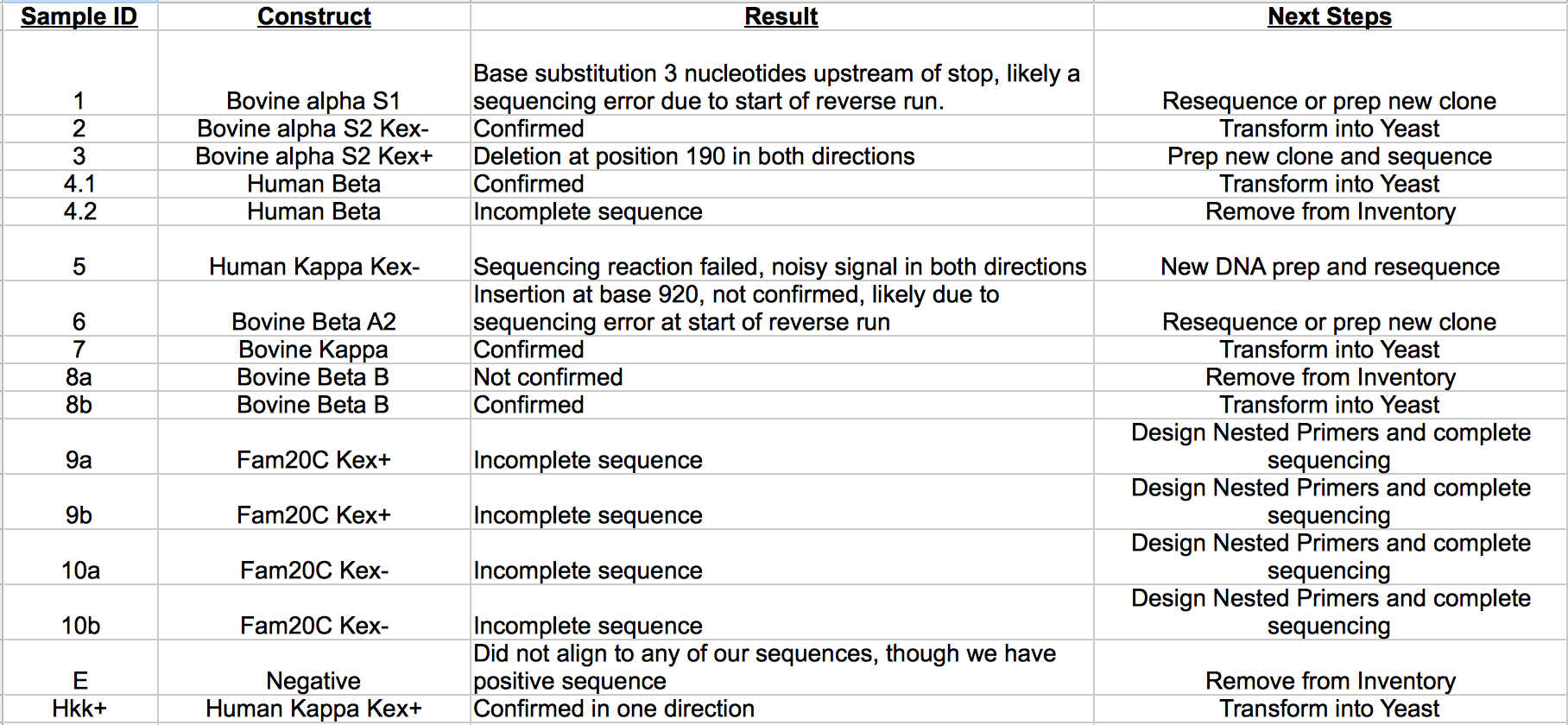
'''Sequencing Analysis Summary Table (October 1, 2014)''' | '''Sequencing Analysis Summary Table (October 1, 2014)''' | ||
[[File:Screen_Shot_2014-10-01_at_4.42.46_PM.png| | [[File:Screen_Shot_2014-10-01_at_4.42.46_PM.png|600px]] | ||
Revision as of 23:46, 1 October 2014
September 30, 2014: DNA pick-up by Sequetech to sequence 16 samples, detailed below
- Location: BioCurious
- Participants: Rachel (set up sample pick-up), Emi (gave samples to Sequetech rep)
- Aims: Confirm correct DNA sequence of our casein and FAM20C inserts in pD1214 vector. Trouble shoot Electra cloning negative control positive colonies (See Cloning bovine beta casein, Fam20C (Kex + & -), from 27Sep2014, "Results and Discussion" September 28, 2014.) Confirm identity of unclearly labeled DNA, prob. hKcasein Kex+ in pD1214.
- Sequencing Strategy & Methods:
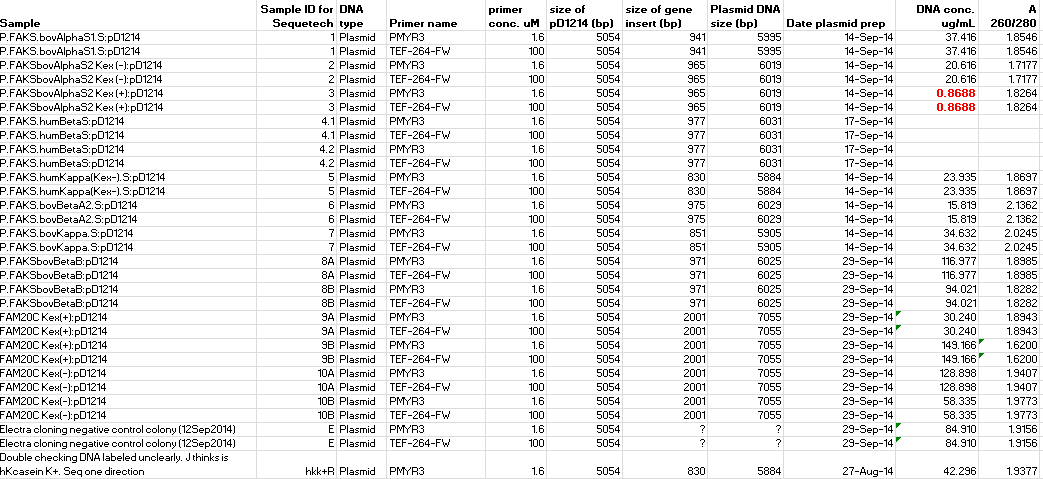
- Submitting 10uL aliquots of 16 samples, detailed in table below, including 10 new casein and FAM20C in pD1214 constructs.
- 4 samples submitted in duplicate -- plasmid preps performed on two separate E. coli clones:
- P.FAKS.humBetaS:pD1214, as previously problematic construct with few transformed colonies (See Troubleshooting failed 12Sep2014 transformation of P.FAKS.humBeta.S for details.)
- P.FAKSbovBetaB:pD121, FAM20C Kex(+):pD1214 and FAM20C Kex(-):pD1214, as plasmid inserts may not be what we expect. (See Cloning bovine beta casein, Fam20C (Kex + & -), from 27Sep2014, "Results and Discussion" September 28, 2014.)
- NOTE: DNA concentration for bovAlphaS2 Kex(+) very low. May have to repeat plasmid prep.
- 4 samples submitted in duplicate -- plasmid preps performed on two separate E. coli clones:
- Sequetech rep. collected samples at ~2:00pm today.
- All except hkk+R to be sequenced in two reactions: the forward (with primer TEF-264-FW) and reverse (with primer PMYR3) directions. Sample hkk+R sampled in single direction as was likely the sample already sequenced in both directions, September 5, 2014.
- TEF-264-FW is a custom primer synthesized through Sequetech: primer sequence recommended by DNA 2.0, suppliers of pD1214. This primer will be maintained in-house for future sequencing reactions. (Note change from sequencing described August 6, 2014, as previous forward primer, Alpha-factor 146-forward, anneals to the FAKS sequence, now inside our redesigned DNA inserts.) See for primer map.
- PMYR3 is an in-house Sequetech primer.
- Submitting 10uL aliquots of 16 samples, detailed in table below, including 10 new casein and FAM20C in pD1214 constructs.
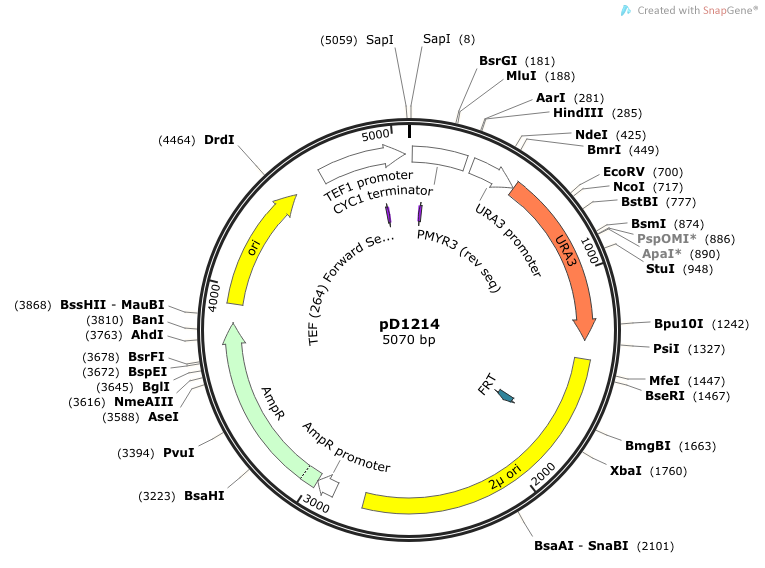
Primer Sequences TEF-264-FW: 5’ TCGATGACCTCCCATTGA 3’ Binds to TEF promoter and reads through in the 5'-3' direction (N-terminus of signal peptide). PMYR3: 5' CTTCCTTTTCGGTTAGAG 3' Binds to the terminator (CYC1) and reads through in the 3'-5' direction (C-terminus of casein).
Location of sequencing primers indicated by small purple arrows on map of pD1214:
Details for 31 sequencing reactions on 16 samples: